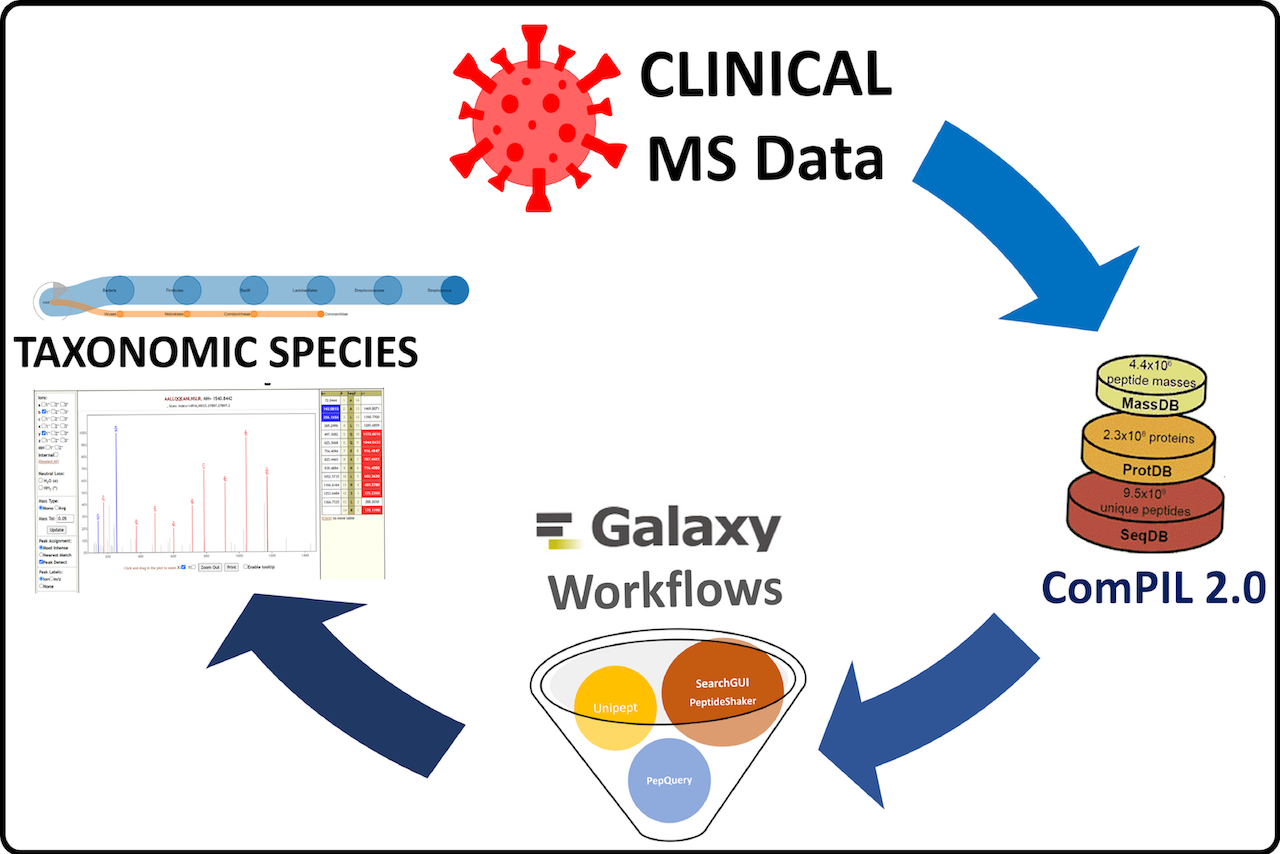
# MS data analysis
Powered by:
Mass spectrometry (MS) based datasets (both from peer-reviewed manuscripts and preprint version) are now available via ProteomeXchange (opens new window) and PRIDE (opens new window). Here, we demonstrate the use of Galaxy workflows to reanalyze MS datasets from cell culture samples (PXD018804, PXD018241, PXD018594), clinical samples (PXD018682, PXD021328, PXD019423, PXD020394, PXD023016,PXD025214, PXD018094, PXD022085) and protein-protein interaction datasets (PXD018117). We also performed metaproteomics analysis on three clinical samples (PXD021328, PXD019423, PXD020394).
- SARS-Cov-2-human-protein-protein interaction (PXD018117 from Krogan Lab)
- Proteomics analysis of SARS-CoV-2 infected cell culture samples (PXD018241 from Matthews lab)
- Proteomics analysis of time-course data from SARS-CoV-2 infected cell culture samples(PXD018594 from Armengaud lab)
- Deep proteomic analysis of COVID-19 virus infected Vero cells. (PXD018804 from Armengaud Lab)
- Proteomics analysis of Gargling samples from CoviD-19 infected patients (PXD019423 from Sinz Lab)
- Proteomics analysis of naso-pharyngeal swabs samples from COVID-19 infected and non-infected individuals (PXD020394 from Lima Lab)
- Proteomics analysis of respiratory tract samples from CoviD-19 infected patients(PXD019119/PXD021328 from Cardozo lab)
- Proteomics analysis of COVID-19 lung tissues(PXD018094 from Zhong lab)
- Proteomic analysis of bronchoalveolar lavage fluid in COVID-19 patients(PXD022085 from Zeng lab)
- Proteomics analysis of naso-pharyngeal swabs samples from COVID-19 infected samples (PXD023016 from Srivastava lab)
- Proteomic analysis of upper respiratory sample COVID-19 patients (PXD025214from Carvalho lab)
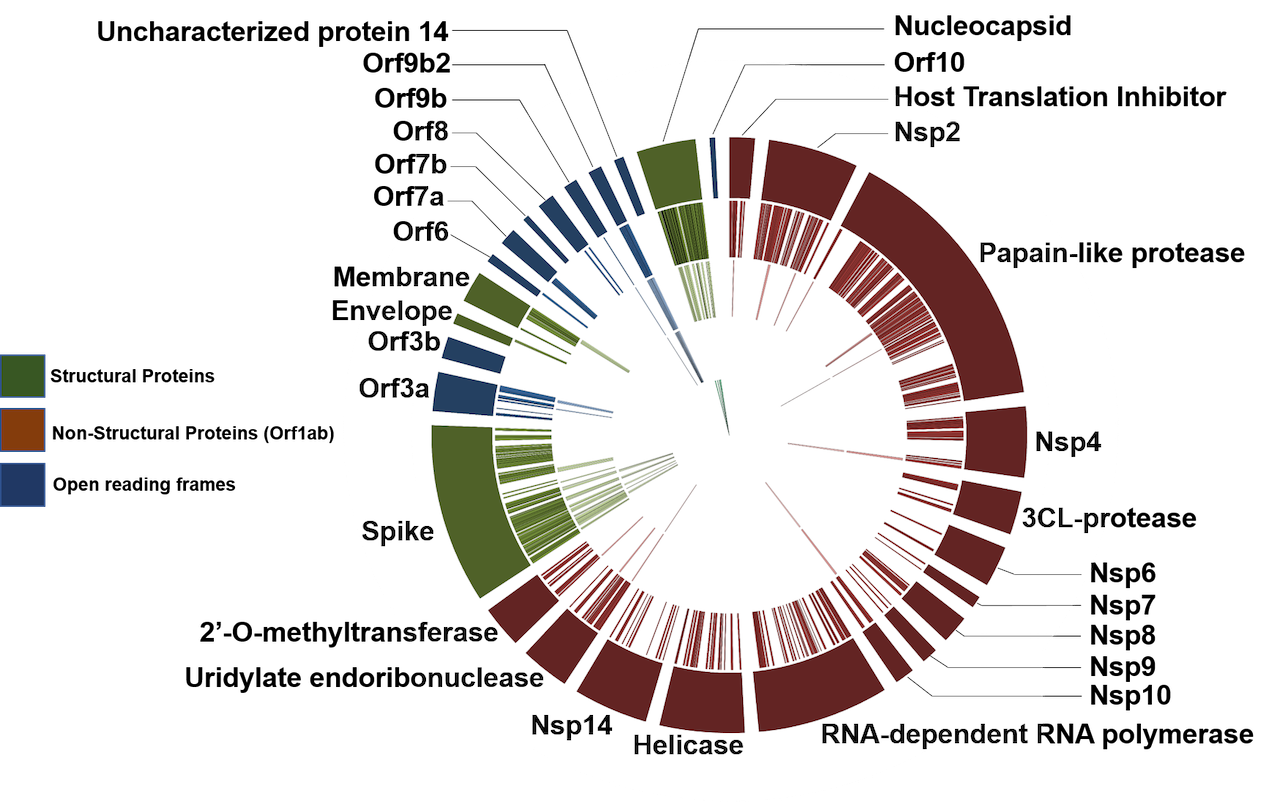
Metaproteomics analysis of Clinical samples now published in JPR (opens new window).
- Metaproteomics analysis of the gargle solutions dataset (PXD019423 from Sinz Lab)
- Reanalysis and metaproteomics analysis of naso-pharyngeal swabs samples from COVID-19 infected and non-infected individuals (PXD020394 from Rivera Lab)
- Reanalaysis and metaproteomics analysis of the respiratory tract samples from CoviD-19 infected patients (PXD021328 from Cardozo Lab)