# Proteomic re-analysis of bronchoalveolar lavage fluid in COVID-19 patients
# Live Resources
| usegalaxy.eu |
|---|
# Description
Zeng lab (opens new window) collected bottom-up mass spectrometry (MS) data on bronchoalveolar lavage fluid in COVID-19 positive patient samples with respiratory failure. Data-dependent acquisition MS spectra were acquired using Q Exactive HF-X mass spectrometer coupled with an EASY-nLC 1200 system and a nano-electrospray ion source.
# Workflow
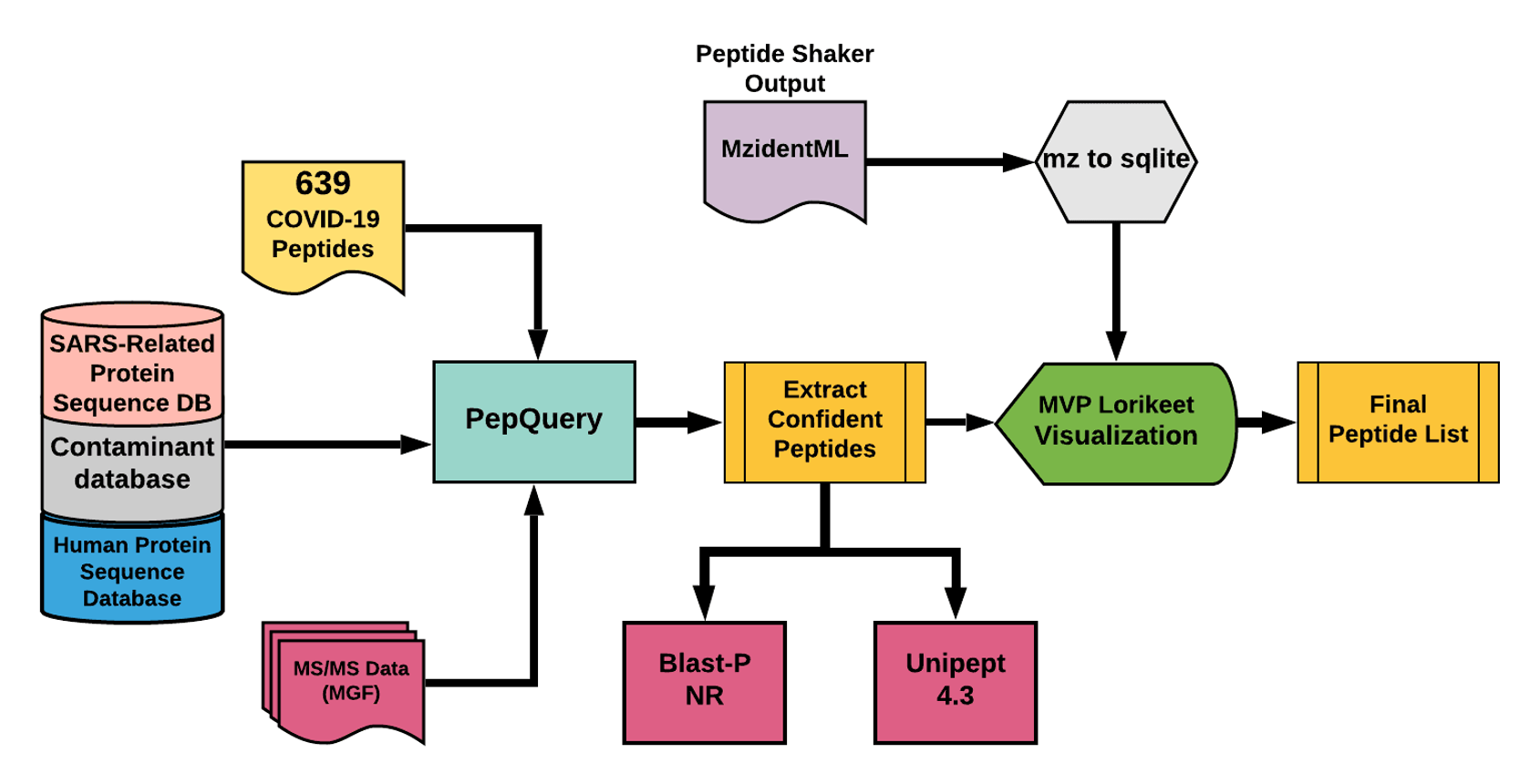
The Galaxy workflow includes RAW data conversion to MGF and mzML format. The MGF files are searched against the combined database of Human Uniprot proteome, contaminant proteins and SARS-Cov-2 proteins database using PepQuery Validation workflow. This resulted in detection of 88 peptides from SARS-CoV-2 proteins. The detected peptides were searched against NCBInr to ascertain that these peptides were specific to SARS-CoV-2 proteins. The detected peptides were later subjected to analysis by Lorikeet visualization to ascertain the quality of peptide identification.
Our Database search workflow did not detect any SARS-CoV2 peptides from both positive and negative samples.
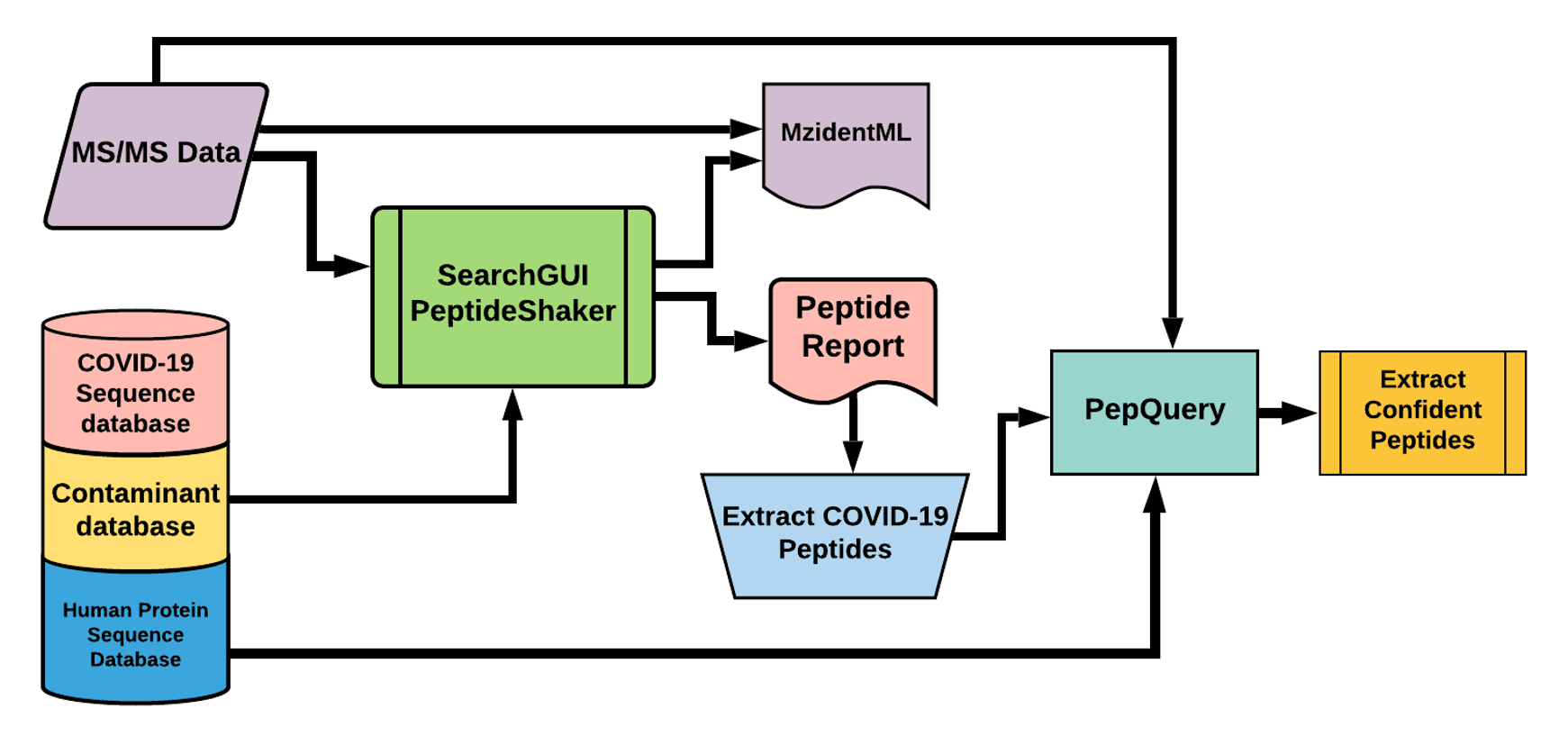
# Results
We detected 36 COVID-19 peptides from all pooled samples in the respiratory tract datasets, We detected 25 SARS-CoV2 peptides from positive patients , 11 SARS-CoV2 peptides from negative patient samples. The peptides were subjected to BLAST-P andLorikeet analysis to ascertain the validity of peptide spectral matches.