# Active site preparation
This section describes the preparation of the active site for docking. The screening of the active site was repeated 17 times, for each of the 17 fragment screening crystal structures that were available at the time (more are expected).
# Live Resources
| usegalaxy.eu |
|---|
# Outline
- A file describing the active site of the protein for each of the fragment screening crystal structures was generated using rDock's
rbcavity(withrbcavity -was -d -r) [1]. - Creating a single hybrid molecule that contains all the ligands - the 'frankenstein' ligand (opens new window).
# History and workflow
A Galaxy workspace (history) containing the most current analysis can be imported from here (opens new window).
The publicly accessible workflow (opens new window) can be downloaded and installed on any Galaxy instance. It contains version information for all tools used in this analysis.
| Active site preparation |
|---|
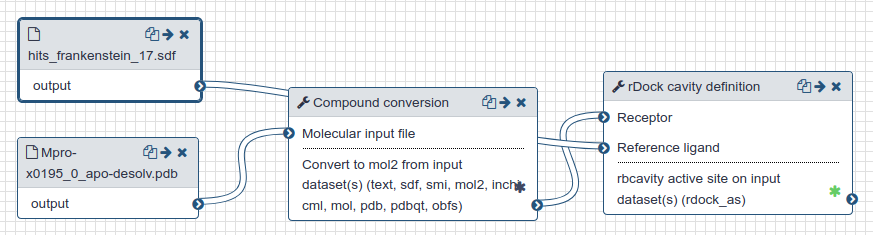 |
| Preparation of the active site for docking. |
# References
[1] Ruiz-Carmona et al. (2014). rDock: a fast, versatile and open source program for docking ligands to proteins and nucleic acids. PLoS Computational Biology. 10 (4). doi:10.1371/journal.pcbi.1003571 (opens new window).